Single and secretly wondering which of your friends might be able to introduce you to your future soulmate?
Sociologists have long studied the dating pool problem, noting that an acquaintance is more likely to introduce you to your mate [1] than a close friend is. Your close friends tend to know all of the same people as you, while acquaintances have greater “bridging social capital,” connecting you to diverse clusters of new people. On the other hand, your close friends might be more motivated to set you up; they care more about your happiness, and they know you better. So who, really, is more likely to set you up? And do these “matchmakers” have special characteristics that make them ideally suited to the task? (Note that when we say “matchmaker” we mean “a friend who introduced you” rather than a professional.)
To answer these questions, a survey was done of approximately 1500 English speakers around the world who had listed a relationship on their profile at least one year ago but no more than two years, asking them how they met their partner and who introduced them (if anyone) and then analyzed network properties of couples and their matchmakers using de-identified, aggregated data.
What does a typical matchmaker looks like?
Unlike the stereotype of an older woman from Fiddler on the Roof, our matchmakers looked pretty similar to the couples they were matching. They were young (most often in their 20s, as were most of the survey respondents), half of them were men, and roughly half of them were single (much like most of the future couple's other friends). So you might not be able to identify your cupid at a glance.
However, our matchmakers had a secret weapon: more unconnected friends. To start, matchmakers have far more friends than the people they're setting up. Before the relationship began, the people who later got matched had an average of 459 friends each, while matchmakers had 73% more friends (794 on average). This isn't that surprising, because network science tells us that your friends usually have more friends than you do [2] . But even compared to the couples' other friends, matchmakers have 8% more friends. So they're unusually well-connected.

Matchmakers have 73% more friends than the typical person they set up.
But more than simple friend count, what matters is whether matchmakers' networks have a different structure. And indeed, they do. Though they have more friends, their networks are less dense: their friends are less likely to know each other. We measure this with the clustering coefficient [3], which is 6% lower for matchmakers than other friends of the couple. Matchmakers also have 4% more isolated nodes as a proportion of their network, or people who have no other mutual friends. Matchmakers typically have about 16 unconnected ties in their networks, and each one is an opportunity: people who both know the matchmaker, but don't yet know each other.

A typical matchmaker's graph has a lot of nodes but relatively few triangles (people with another friend in common). Each pair of unconnected nodes is an opportunity for an introduction.
Are matchmakers close friends or acquaintances?
Matchmakers were pretty close friends to the people they later introduced. (Makes you wonder why they didn't introduce their friends sooner!) More than half of the survey respondents said they felt “very” or “extremely” close to the matchmaker before the match was made. Nearly half spoke to the matchmaker every day, and two thirds spoke to their matchmaker more than once a week. And half had known the matchmaker for three years or more, as did their future partner. So by every measure, matchmakers were close friends with the future couple.

Matchmakers were more likely to be close friends, rather than acquaintances.
However, survey responses are clouded by retrospective bias. Wouldn't you grow closer over time to the person who introduced you to your partner? And that rosy haze can make it hard to remember how close you really felt two years ago. So, for that an examination was done on the network data from the time before the relationship to see if people were actually close to their matchmaker and then counted the number of mutual friends each person in the couple had with the matchmaker before the relationship started, a common proxy for relationship closeness. Matchmakers typically had 57 friends in common with either person in the future couple, amounting to about 6% of their pool of friends. Does that make them close friends? One good comparison point is the couple today: they typically have 52 friends in common, or about 7% of their pool of friends. If romantic partners are your gold standard for close friends, the matchmakers look like strong ties!
Notably, the fraction of mutual friends between the matchmaker and a member of the couple is significantly higher when they are of the same gender, around 7.4% when the relationship started compared to only 5.1% when they are of the opposite gender.
Couples who met through a friend vs. those who met another way: How do their networks differ?
Couples who were introduced by a friend are more likely to have at least one friend in common a year prior to the start of their relationship (84% of them) compared to those who met in other ways (74% only), which is to be expected – this quantity increases over time, and a few months into the relationship, more than 99% of couples have a friend in common, regardless of how they met.
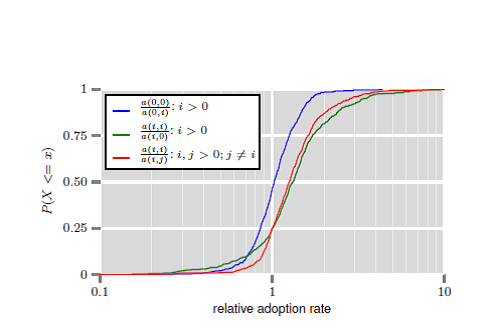
More interesting though, is that the fraction of mutual friends starts higher for couples who were *not* introduced by a mutual friend (around 4.4% overlap 18 months prior to the start of the relationship) compared to those who were (about 3.4% at that time). It is notable that that quantity increases faster for the latter, to the point that 18 months into the relationship the overlap of social networks of both members of the couple averages at around 7.7% whether or not they were introduced by a friend.
We can understand this by the fact that matchmakers act as bridges between communities or social groups: when you meet someone at school or at work, both you and the other person evolve in the same social circles, hence you have a higher overlap. But imagine one of your friends introduced you to someone at a different school; you might have fewer friends in common with that person at first, but over time you both start making new friends in each other's schools.

Couples who were introduced by a friend had a smaller fraction of mutual friends prior to the relationship.
If no one introduced them, how did people meet their partner?
Couples who found each other without a friend introducing them most often met at school. But nearly 1 out of 5 couples met through an online dating service or app. Keep in mind these are primarily couples in their twenties and thirties who started dating in 2013 or early 2014.
Fun fact: As many couples met through Facebook as at a party or in a bar.
This is consistent with a Pew study [4] finding that many social network site users have followed or friended someone because one of their friends suggested they'd want to date that person. And many of our respondents cited Facebook as playing a role in the early stages of their relationships: (Quotes have been edited for length and names have been changed.)
Dan first became interested in me because of a picture he saw of me on facebook. I met the person who introduced me to Dan at a party that was organized on Facebook. Now that I think of it, it's possible that I wouldn't even know Dan if Facebook didn't exist.
We were acquaintances, but after she posted on my wall that we had a lot in common, we began dating.
Without that friend suggestion in his newsfeed, I am not sure he and I would be together today!

People most often met their partner at school. As many couples met through Facebook as met at a party or bar.
References
1. Interpersonal Ties (http://en.wikipedia.org/wiki/Interpersonal_ties)
2. Friendship Paradox (http://en.wikipedia.org/wiki/Friendship_paradox)
3. Clustering Coefficient (http://en.wikipedia.org/wiki/Clustering_coefficient#Local_clustering_coefficient)
4. Pew Study (http://www.pewinternet.org/files/old-media//Files/Questionnaire/2013/Survey%20Questions_Online%20Dating.pdf)